DeepMind’s AI for protein structure is coming to the masses
You can also search for this author in PubMed Google Scholar
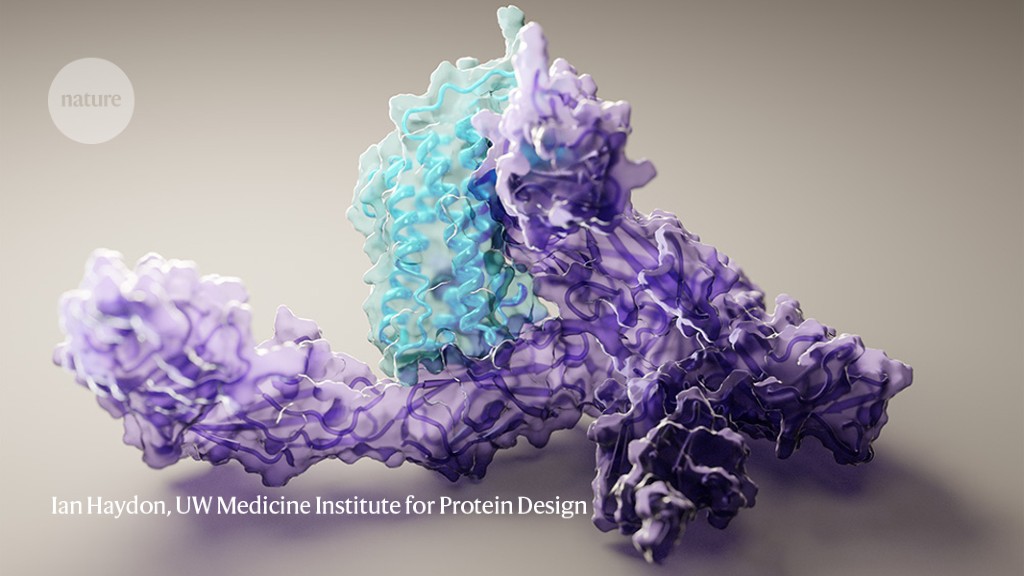
The structure of human interleukin-12 protein bound to its receptor, as predicted by machine-learning software. Credit: Ian Haydon, UW Medicine Institute for Protein Design
It’s protein-structure prediction for the people. Software that accurately determines the 3D shape of proteins is set to become widely available to scientists.
On 15 July, the London-based company DeepMind released an open-source version of its deep-learning neural network AlphaFold 2 and described its approach in a paper in Nature1. The network dominated a protein-structure prediction competition last year.
Meanwhile, an academic team has developed its own protein-prediction tool inspired by AlphaFold 2, which is already gaining popularity with scientists. That system, called RoseTTaFold, performs nearly as well as AlphaFold 2, and is described in a paper in Science paper also published on 15 July[2]2.
The open-source nature of the tools means that the scientific community should be able to build on the advances to create even more powerful and useful software, says Jinbo Xu, a computational biologist at the University of Chicago in Illinois, who was not involved in either effort.