Engineering Plastic-Degrading Enzymes and PCSK9 Binders with Protein AI Tools
In the past year, a new class of machine learning tools for protein engineering promise to change how we develop drugs and enzymes. However, in these early days, the industry is still trying to understand how to use these tools to do concrete things.
After working with cutting-edge academic labs and computational biotech teams on real problems in research and industry, we’ve assembled a curated toolbox of graphically accessible protein engineering tools and demonstrate how to string them together for useful molecular design tasks.
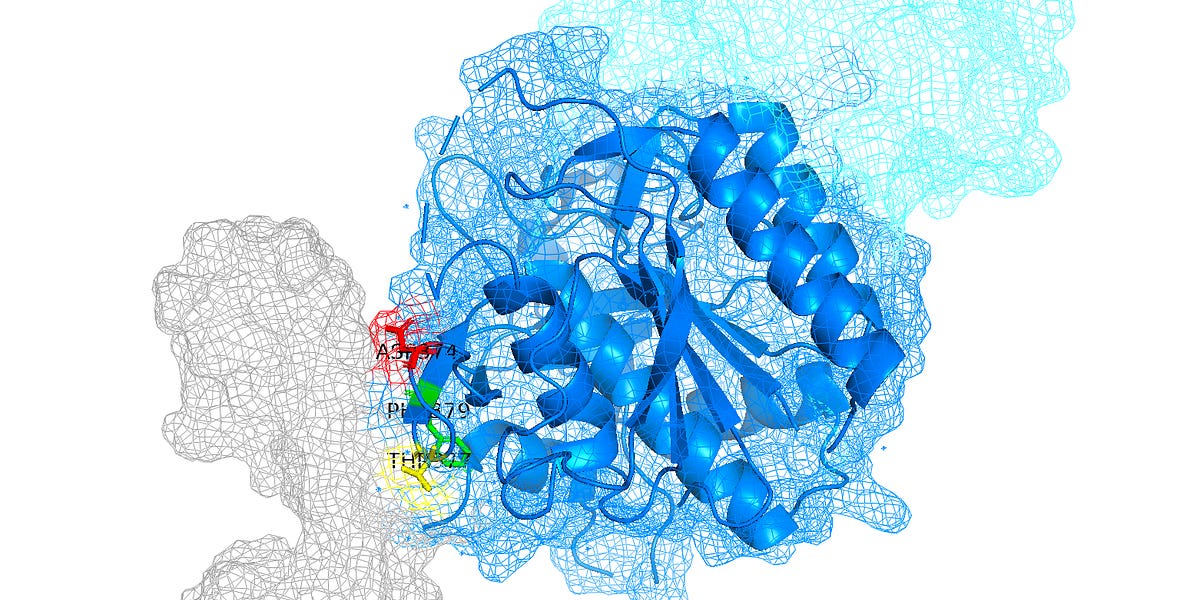
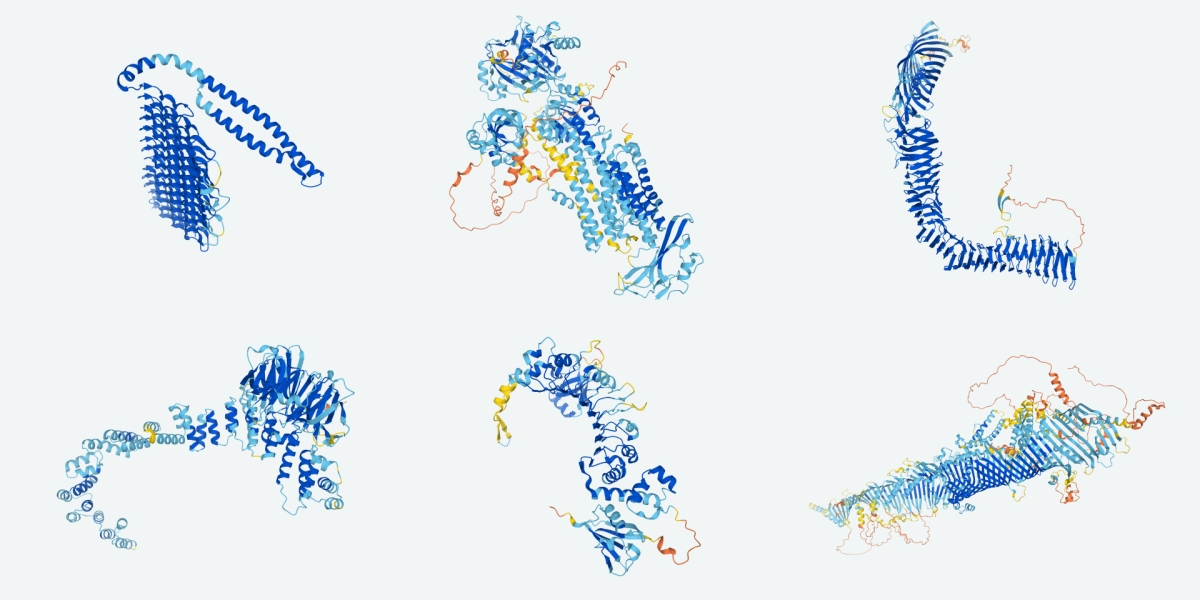
Here we build two different proteins - plastic-degrading enzymes and blood cholesterol drugs - each with distinct scientific goals to show the flexibility of the toolkit and breadth of utility. To build enzymes we: 1. Finetune ProtGPT2 on enzyme sequences 2. Use finetuned ProtGPT2 to generate a library of novel enzymes 3. Create 3D protein structures with OmegaFold 4. Predict thermal stability with TemStaPro 5. Calculate electrostatic potential and hydrophobicity with PEP-Patch 6. Evaluate aggregation propensity with Aggrescan3D To build PCSK9 binders we: 1. Use Pymol to identify the region of PCSK9 we want to bind 2. Diffuse 10 scaffolds near binding hotspots on PCSK9 with RFDiffusion 3. Use ProteinMPNN to generate 100 sequences for these scaffolds 4. Predict 3D protein structure with ColabFold
We trace the actual end-to-end engineering flows from ideation to structure and use believable scientific details to contextualize the problems in familiar biology. We hope this post serves as a resource for scientists and engineers navigating this new ecosystem of models.












/cdn.vox-cdn.com/uploads/chorus_asset/file/25686563/ploopy1.jpg)